Bing Yang Lab @Purdue
Liquid-liquid phase separation (LLPS) is a biophysical process where a homogeneous solution of molecules (typically proteins, nucleic acids, or both) separates into two distinct liquid phases—one dense and one dilute—much like oil separating from water. It is postulated that LLPS helps compartmentalize cellular processes without membranes, regulating activities like RNA metabolism, signal transduction, stress response, and gene expression. And abnormal phase separation has been associated with pathological aggregates implicated in diseases such as ALS, Alzheimer's, and certain cancers.
Recently, we found that the Microprocessor complex forms biomolecular condensates in select tissues in C. elegans. Our future research aims to further understand the composition and function of the Microprocessor condensates in multiple species. We will characterize the Microprocessor condensates in both animal model and human iPSCs to investigate the molecular mechanics that underpin condensate formation, dynamics, and function in native cells. We are also interested in exploring potential molecular condensates formed by other miRNA biogenesis machineries.
DRSH-1::GFP granules in C. elegans hypodermis cells
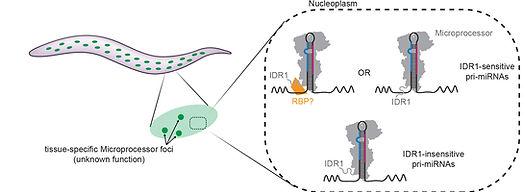
Precise control of miRNA biogenesis is of extreme importance, since mis-regulation of miRNAs underlies or exacerbates many disease states. Further research is necessary to understand how intrinsically disordered regions (IDRs) in essential miRNA biogenesis machineries function, as these regions remain largely uncharacterized. Recent studies from different groups including ours have shown that IDR region of Drosha is critical for the biogenesis of a subset of microRNAs through the cis elements in the primary miRNA transcript. We are trying to understand the precise contributions of individual structural and sequence features of pri-miRNAs to their processing efficiency and accuracy.
Publications
B. Yang, B. Galletta, R. Sakhawala, N. Rusan, K. McJunkin. (2025) An intrinsically disordered region of Drosha selectively promotes miRNA biogenesis, independent of tissue-specific Microprocessor condensates. Genes & Development. In Press.
K. Kotagama, A. L Grimme, L. Braviner, B. Yang, R. Sakhawal, G. Yu, L. K Benner, L. Joshua-Tor, K. McJunkin. (2024) Catalytic residues of microRNA Argonautes play a modest role inmicroRNA star strand destabilization in C. elegans. Nucleic Acid Research DOI:10.1093/nar/gkae170.
Y. Li, X. Dou, J. Liu, Y. Xiao, Z. Zhang, L. Hayes, R. Wu, X. Fu, Y. Ye, B. Yang, LW Ostrow, C.He, S. Sun. (2023) Globally reduced N6-methyladenosine (m6A) in C9ORF72-ALS/FTDdysregulates RNA metabolism and contributes to neurodegeneration. Nature Neuroscience26(8):1328-1338.
BJ Galletta, R. Varadarajan, C. J Fagerstrom, B. Yang, K. Haase, K. McJunkin, NM Rusan.(2023) The E3 ligase Poe promotes Pericentrin degradation. Molecular Biology of the Cell34(9):br15.
B. Donnelly, B. Yang, A. Grimme, K-F. Vieux, C-Y. Liu, L. Zhou, K. McJunkin. (2022) Thedevelopmentally-timed decay of an essential microRNA family is seed sequence-dependent.Cell Reports 40(6):111154.
B. Yang, M. Schwartz, K. McJunkin. (2020) In vivo CRISPR screening for phenotypic targets ofthe mir-35-42 family in C. elegans. Genes & Development 34:1227-1238.
B. Yang, K. McJunkin. (2020) CRISPR screening strategies for microRNA target identification.FEBS J 287:2914-2922.
B. Yang, K. McJunkin. (2020) The mir-35-42 binding site in the nhl-2 3’UTR is dispensable fordevelopment and fecundity. Micropublication Biology.
B. Yang, X. Xu, L. Russell, M. Sullenberger, J. Yanowitz and E. M. Maine. (2019) A DNA repairprotein and histone methyltransferase interact to promote genome stability in theCaenorhabditis elegans germ line. PLOS Genetics 15(2):e1007992.
B. Mutlu, H-M Chen, J.J. Moresco, B.D. Orelo, B. Yang, J. Gaspar, S. Keppler-Ross, J.R. YatesIII, D.H. Hall, E.M. Maine, and S.E. Mango. (2018) Regulated nuclear accumulation of a histonemethyltransferase times the onset of heterochromatin formation in C. elegans embryos.Science Advances 4(8): eaat6224.
J. L. Lissemore, E. Connors, Y. Liu, L. Qiao, B. Yang, M. L. Edgley, S. Flibotte, J. Taylor, V. Au,D. G. Moerman, and E. M. Maine. (2018) The molecular chaperone HSP90 promotes Notchsignaling in the germline of Caenorhabditis elegans. G3 doi: 10.1534/g3.118.300551.
Y. Guo, B. Yang, Y. Li, X. Xu and E. M. Maine. (2015) Enrichment of H3K9me2 on unsynapsedchromatin in Caenorhabditis elegans does not target de novo sites. G3 5(9): 1865-1878.